 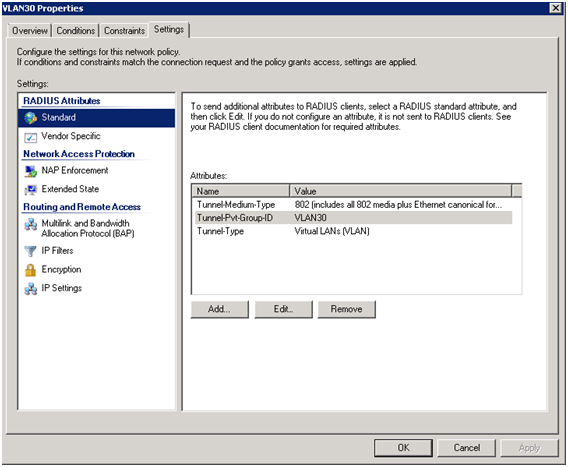
The disadvantage of a high value is that this effectively reduces the size of the training set.Ī low value (like 150) means that all training sequences are taken into account, except The advantage is that the training is more specific to the target GC content. (default: 200) determine the influence of the deviation in GC content.Ī high value (like 300) means that mostly training sequences with a very similar GCĬontent to the target GC content are taken into account. The two non-zero numbers in the middle of the 4x4 Matrix in bug_weightmatrix.txt It gets a weight that is the higher, theĬloser the GC content of the training sequence is to the target GC content. Given a target GC content, each sequence in the training set is weighted depending on how similar its GC content These parameters can be set in the meta parameters file. Constant/gc_range_min and /Constant/gc_range_max. AUGUSTUS uses /Constant/decomp_num_stepsĭifferent sets of parameters, each for a different GC content. 
Constant/decomp_num_steps is the number of different levels of GCĬontent that is taken into account, i.e. 
I describe here only the dependency on the GC content. Parameters depend on the average frequencies of the 4 bases in the query sequence.įor example, in human, the GC content staysĬonsistently above or below average over long sequence stretches (isochores).ĪUGUSTUS can locally use different parameters that are adjusted to the base composition This section describes the meaning of the meta parameter /Constant/decomp_num_stepsĪnd the matrix in the file bug_weightmatrix.txt and you can savely skip thisįor some species like humand and honey bee it makes sense to let the model The meta parametersĭetermine how the parameters are calculated.Ĭreate the files for training " bug" from a template. There are a few dozen meta parameters but many thousands of parameters. Splice site patterns, the k-mer probabilities of coding and noncoding regions. We call parameters like the size of the window of the splice site models and the order of the Markov model meta parameters, in contrast to parameters like the distribution of CREATE A META PARAMETERS FILE FOR YOUR SPECIES 40 letters + gt + 40 lettersĪss: donor (=3') splice site. Randomly split the set of annotated sequences in a training and a test set.ĭss gccgagaactccgctcgttctgtgcgttctcctgtcccaggtagggaagaggggctgccgggcgcgctctgcgccccgtttcĭss cgtgattgtcggggggaaagacatccagggctccttgcaggtaacacatctgtttgagataacttgggttcaaggaggacatĭss agagaatcagagacagcctttcccaagagatgttggcaaggtaagtcagacaaacagcaaatgacaaaaacatgtttttatgĭss cattgtcactgttgtgtcacctgcgctgctggaccgagaggtgagctgaaaagaataccactttctttttcacgagaatagaĭss tgacaaaaatgatcactcaccaaaattcaccaagaaagaggtaaacccctgtgccaaacaccaaccaccactgtggtcacagĪss gttagtatgcttctttaattttttttctccctgaaattataggaaccagatgttaaaaaattagaagaccaacttcaaggcgĪss -ggctttgtctttgcagaatttatagagcggcagcacgcaaagaacaggtattactaĪss gattccttgtgattagcctctcttgctccttttctccaccagcaaagtcgaccaagaaattatcaacattatgcaggatcggĪss aaccgtagtaaacagcatgaatcgtgttttgtttttgaacagaccactggccttgtgggattggctgtgtgcaatactcctcĭss: donor (=5') splice site. the CEGMA pipeline to identify the structure of core eukaryotic genesġ.2 Split gene structure set into training and test set.pompe, but it may fail on very complex genomes. This was tested to work very well on Drosophila, C. Both programs are automatically trained and genes are predicted genome-wide using the RNA-Seq. It uses a genome and RNA-Seq alignments as input. BRAKER: a new pipeline that combines GeneMark-ET and AUGUSTUS.Iteration of training with predicted genes, starting with an existing parameter set.This approach is described in section Using Scipio to create a training set. spliced alignments of protein sequences of the same or a very closely related species against theĪssembled genomic sequence, e.g.spliced alignments of de novo assembled transcriptome short reads (RNA-Seq).spliced alignments of ESTs against the assembled genomic sequence (e.g.1.1 Options for compiling a set of gene structures For the exact format that the training program can read in lookĪs an example at one of the training genbank files at the augustus web server: Store the sequences together with their annotation Not overlap, and only one transcript per gene is allowed. This is of course also very important when you want to test the accuracy on a test set.Įach sequence can contain one or more genes, the genes can be on either strand. The non-redundancy is very important to avoid overfitting during optimize_. 
My criterion is: No two genes in the set are more than 80% identical on the amino acid level. If in two different sequences genes withĪn almost identical amino acid sequence is annotated then delete one of them. Non-redundancy is important if you plan to use part of it for training and another partįor assessing prediction accuracy. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
